High Content Genomics and Bioinformatics
The facility has several computer workstations dedicated to data analysis and makes use of the high performance computing facility hosted at the IGTP data processing center. Besides our in house laboratory infrastructure to carry on sample quality control and preparation for high content genomic analysis assays the unit is coordinated with centralized genomics core facilities such as the CRG-CNAG.
Documents
Library quality control request
Micro-array hybridisation request
Next generation sequencing request
Bioinformatics analysis request
Robotic liquid handling

Sciclone NGS (PerkinElmer) liquid handling robot
Robot with 96 disposable tip head and plate gripper. Protocols for automated execution of molecular biology applications, especially for next generation sequencing (NGS). Allows parallel handling of up to 96 samples simultaneously. Used for NGS library preparation and quantification and 384 microwell plate qPCR setup.

Evo-150 (Tecan) liquid handling robot
The robot is equipped with 8 fixed tips and set up for automated sample processing for Illumina infinium assays. It allows parallel handling of up to 384 samples.
Nucleic acid quantification
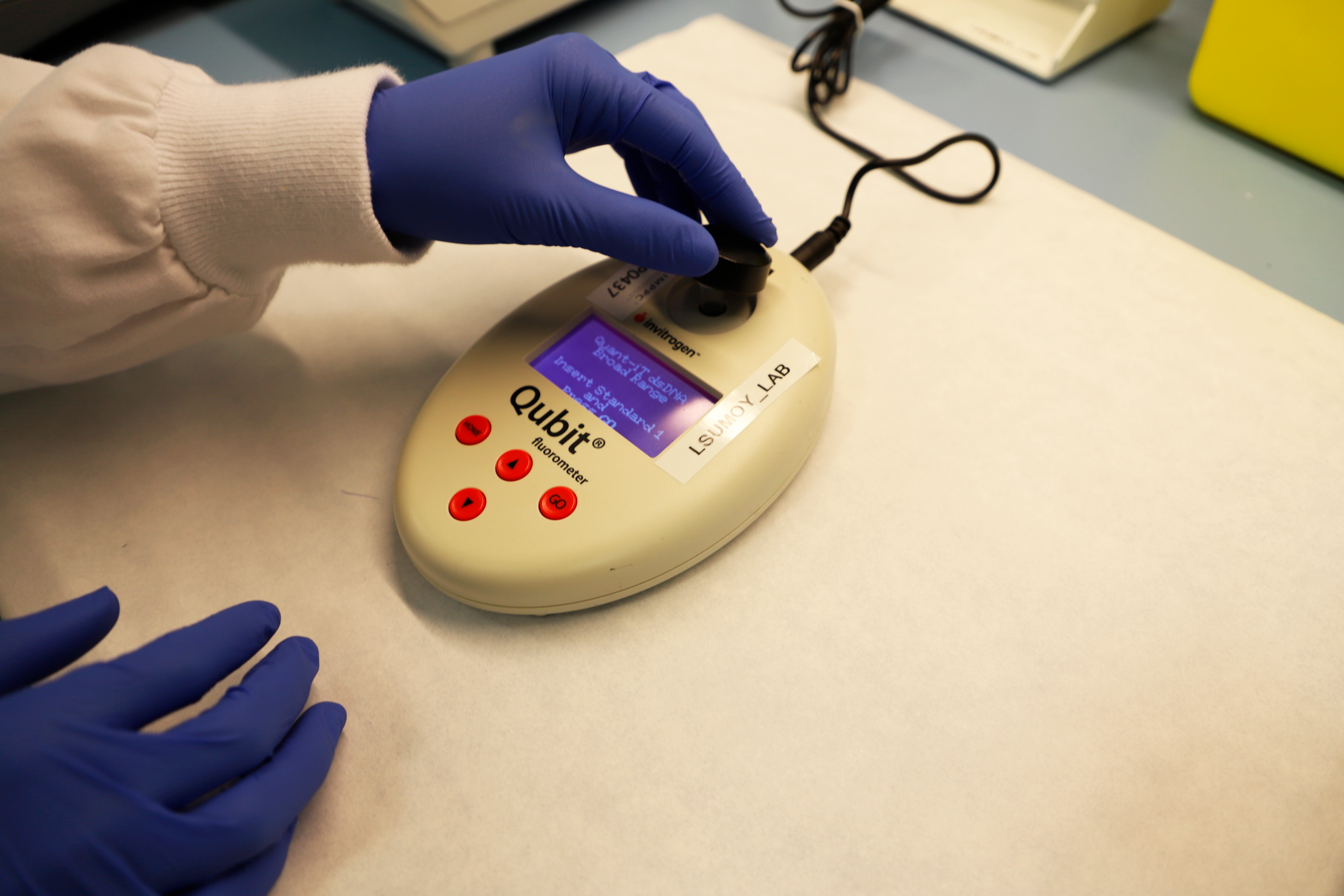
Qubit (Thermofisher)
Single tube fluorometer. Used for precise nucleic acid quantification from 1-20 µl of sample. Discriminates single from double stranded DNA, and from RNA.
Bioinformatic data analysis computing

HP xw8600, xw4600, Z600, Z440 (Hewlet Packard) workstations
High performance computing workstations with Linux operating system. Used for local computation or remote access to centralized nodes and storage systems hosted at the IGTP scientific computing data processing center (HPC) for bioinformatic analysis of genomic data generated by arrays and NGS.

Dell Precision T7500 (Dell)
High performance computing server with Linux operating system to support next generation sequencing data and other memory intensive data analysis.
Automated nucleic acid analysis and size selection

Bioanalyzer 2100 (Agilent)
Automated lab-on-a-chip microfluidic chip nano-electrophoresis system. Used for analysis of DNA and RNA integrity and size assessment, and smallRNA qualitative profiling. Allows procesing of up to 11-12 samples per run.

Pippin Prep (SAGE)
Automated gel electrophoresis and elution system. Used for size selection of DNA libraries in next generation sequencing applications libraries. Allows simultaneous separation of up to 4 library pools per run and predefined size fraction elution using independent settings per pool.
