A new study with the GCAT cohort analyses pigmentary traits and their relationship with diseases

Researchers from the GCAT|Genomes for Life project of the IGTP have analysed the genetic basis of 13 pigment traits and their association with diseases. The work has been published this month in Genes.
Pigmentary traits, genetically determined, have been largely studied by GWAS mostly in Caucasian ancestry cohorts from North Europe. However, a lack of information is remarkable in South European populations. Understanding the genetic basis of adaptation of skin colour in different populations has many implications not only for human evolution, but also for medicine practice.
Using the GCAT cohort, Xavier Farré and Natalia Blay, the two first authors of the study, together with Betty Cortés, have built a comprehensive analysis strategy leading to the identification of phototype associated-diseases in a total of nearly 20,000 Catalan subjects and exploring 177 ICD-9 diagnoses. In addition, genome-wide analysis of more than 10M variants, including structural variants across pigmentary traits, using polygenic risk scores, identified two new regulatory genes involved in skin colour and revealed shared genetics among diseases and pigmentation determinants.
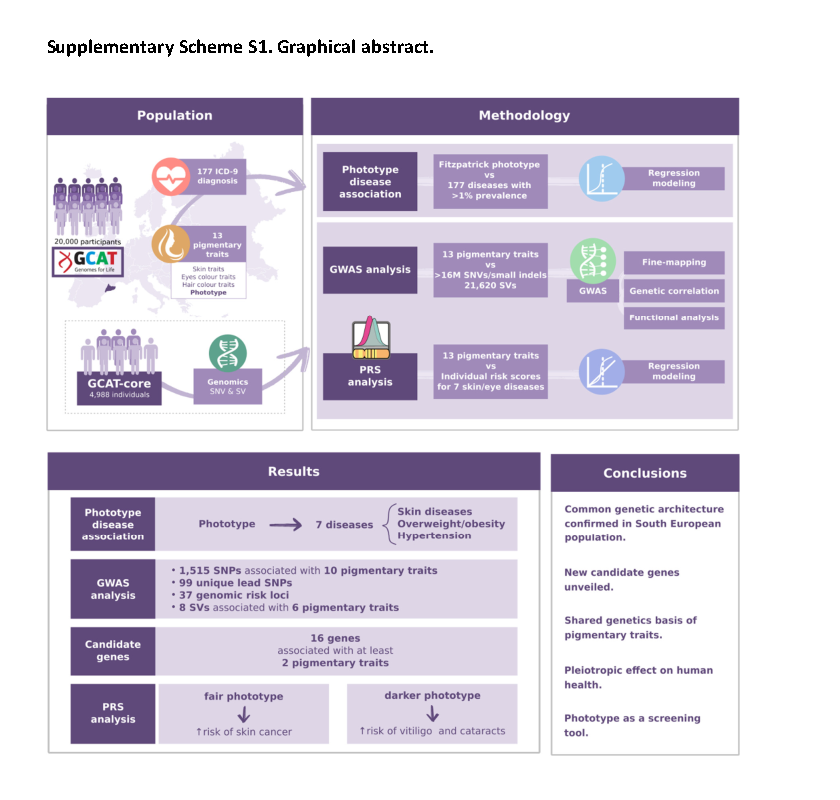
“We observed that a fair skin phototype is strongly associated with non-melanoma skin cancer and other dermatoses and confirmed by PRS-approach the shared genetic basis with some skin and eye diseases, such as melanoma, non-melanoma skin cancer, basal cell carcinoma. On another note, a darker phototype associated with vitiligo and cataracts. The study also identified a link with obesity and hypertension” explains Rafael de Cid, senior author of the study.
Genetic analyses revealed 37 risk loci associated with most pigmentary traits, from which 16 genes significantly associated with at least two pigmentary traits. Some of them have been widely reported, such as MC1R, HERC2, OCA2, TYR, TYRP1, SLC45A2, and three new candidates (C1QTNF3, C1QTNF3-AMACR, and LINC02876) with regulatory potential were unveiled: a long non-coding RNA (LINC02876) and non-sense mediated decay mRNA (NMD)(C1QTNF3-AMACR), defining a putative complex hotspot locus for the fine-regulation of the pigmentary traits.
This study unveils the genetic basis of pigmentary traits in the GCAT cohort, being the largest South European population analysed so far, extending previous work conducted in mostly in Caucasian ancestry cohorts from North Europe, and including a broader spectrum of diseases than previous studies. "Our results highlight the importance of incorporating population information to adapt personalized care approaches to manage differential risk in general population, being of great importance in modern mixed populations. Skin color genetics, as a proxy of skin functionality, could help disease screening in mixed populations", says de Cid.
Further research taking on multi-ancestry cohorts will be able to draw definitive conclusions about the association between pigmentary characteristics and the prevalence of diseases in different populations.
Reference
Farré X, Blay N, Cortés B, Carreras A, Iraola-Guzmán S, de Cid R. Skin Phototype and Disease: A Comprehensive Genetic Approach to Pigmentary Traits Pleiotropy Using PRS in the GCAT Cohort. Genes (Basel). 2023 Jan 5;14(1):149. DOI: 10.3390/genes14010149. PMID: 36672889; PMCID: PMC9859115.
